Histopathology slide is the gold standard of cancer diagnosis. It delivers massive information on tumor microenvironment (TME), which not only plays a vital role in interpreting tumor initiation and progression, but also influences therapeutic effect and prognosis of cancer patients. The crosstalk between different types of tissues are highly related to tumor progression. Therefore, it is urgent to segment and differentiate different tissues for further clinical researches.
Lung cancer is the leading cause of cancer death worldwide . In this challenge, we aim to perform tissue semantic segmentation in H&E stained Whole Slide Image (WSI) for lung adenocarcinoma. The current challenge is that obtaining pixel-level annotations of tissue semantic segmentation is extremely difficult and time-consuming. Inspired by Weakly Supervised Semantic Segmentation (WSSS) in computer vision, we decided to provide only image-level annotations to perform tissue semantic segmentation.
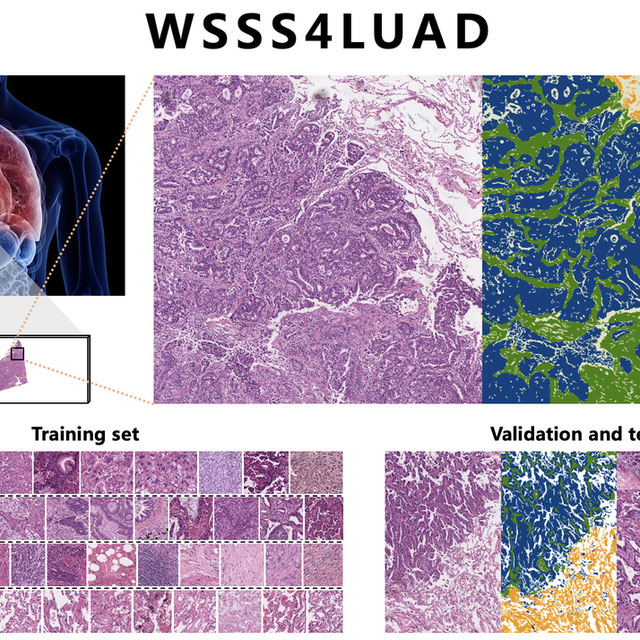
In this challenge, we scanned 67 H&E stained slides from Guangdong Provincial People’ Hospital (GDPH) and collected 20 WSIs from The Cancer Genome Atlas (TCGA). Only one WSI was extracted per patient. The goal of this challenge is to use only image-level annotations to achieve pixel-level prediction of three common and meaningful tissue types, tumor epithelial tissue, tumor stromal tissue and normal tissue. Participants are only given image-level annotations (3-digit one-hot encoding) for machine learning algorithm training, and pixel-level ground truth for validation and testing.
Awards:-
Top 3 ranking winners will receive 500$, 400$, 300$, respectively. To be eligible for the price, the source code and the model parameters must be submitted, besides the results.
Deadline:- 30-09-2021